  | Grentzinger, V., Artesi, M., Gonon Rodrigues Palmeira, L., Dideberg, V., Olivier, S., Bours, V., & Durkin, K. (24 April 2024). Hidden in Plain Sight: Homologous Recombination masks Filaggrin Alleles of Potential Clinical Significance [Poster presentation]. byteMAL 2024, Maastricht, Netherlands. |
  | Tang, L., Swedlund, B., Dupont, S., Harland, C., Costa Monteiro Moreira, G., Durkin, K., Artesi, M., Mullaart, E., Sartelet, A., Karim, L., Coppieters, W., Georges, M., & Charlier, C. (09 March 2024). GWAS reveals determinants of mobilization rate and dynamics of an active endogenous retrovirus of cattle. Nature Communications, 15 (1), 2154. doi:10.1038/s41467-024-46434-1  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
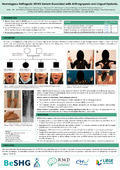  | Mouraux, C., FOUQUET, C., Artesi, M., Durkin, K., BULK, S., & Depierreux, F. (2024). Homozygous pathogenic MYH3 variant associated with arthrogryposis and lingual dystonia [Poster presentation]. BeSHG annual symposium 2024, Belgium.  Peer reviewed Peer reviewed |
  | Grentzinger, V., Durkin, K. W., Gonon Rodrigues Palmeira, L., Artesi, M., Olivier, S., & Bours, V. (24 November 2023). Identification of novel haplotypes within the challenging medically relevant FLG gene [Poster presentation]. SFMBBM PhD Day, Liège, Belgium. |
  | Rudin, C., Bollen, N., Hong, S. L., Wegner, F., Politi, L., Mellou, K., Geenen, C., Gorissen, S., Verhasselt, B., Durkin, K., Henin, C., Logist, A.-S., Dellicour, S., Resa, T., Stadler, T., Maes, P., Cuypers, L., André, E., Egli, A., & Baele, G. (November 2023). Investigation of an international water polo tournament in Czechia as a potential source for early introduction of the SARS-CoV-2 Omicron variant into Belgium, Switzerland and Germany, November 2021. Euro Surveillance: Bulletin Européen sur les Maladies Transmissibles, 28 (45). doi:10.2807/1560-7917.ES.2023.28.45.2300018  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Kremer, C., Torneri, A., Libin, P. J. K., Meex, C., Hayette, M.-P., Bontems, S., Durkin, K. W., Artesi, M., Bours, V., Lemey, P., Darcis, G., Hens, N., & Meuris, C. (16 June 2023). Reconstruction of SARS-CoV-2 outbreaks in a primary school using epidemiological and genomic data. Epidemics, 44, 100701. doi:10.1016/j.epidem.2023.100701  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Grentzinger, V., GRISART, B., Meynsbrughen, M., Artesi, M., Gonon Rodrigues Palmeira, L., Dideberg, V., CHARLOTEAUX, B., Durkin, K. W., Brysse, A., & Bours, V. (25 May 2023). Detecting pathogenic variants in chronic kidney disease using long-read sequencing [Paper presentation]. Annual Meeting @ Living Tomorrow Vilvoorde, Vilvoorde, Belgium. |
  | Cuypers, L., Keyaerts, E., Hong, S. L., Gorissen, S., Menezes, S. M., Starick, M., Van Elslande, J., Weemaes, M., Wawina-Bokalanga, T., Marti-Carreras, J., Vanmechelen, B., Van Holm, B., Bloemen, M., Dogne, J.-M., Dufrasne, F., Durkin, K., Ruelle, J., De Mendonca, R., Wollants, E., ... Van Weyenbergh, J. (22 May 2023). Immunovirological and environmental screening reveals actionable risk factors for fatal COVID-19 during post-vaccination nursing home outbreaks. Nature Aging, 3 (6), 722 - 733. doi:10.1038/s43587-023-00421-1  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Cuypers, L., Dellicour, S., Hong, S. L., Potter, B. I., Verhasselt, B., Vereecke, N., Lambrechts, L., Durkin, K., Bours, V., Klamer, S., Bayon-Vicente, G., Vael, C., Ariën, K. K., De Mendonca, R., Soetens, O., Michel, C., Bearzatto, B., Naesens, R., Gras, J., ... Baele, G. (20 October 2022). Two Years of Genomic Surveillance in Belgium during the SARS-CoV-2 Pandemic to Attain Country-Wide Coverage and Monitor the Introduction and Spread of Emerging Variants. Viruses, 14 (10), 2301. doi:10.3390/v14102301  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | El Moussaoui, M., Maes, N., Hong, S. L., Lambert, N., Gofflot, S., DELLOT, P., Belhadj, Y., Huynen, P., Hayette, M.-P., Meex, C., Bontems, S., Defêche, J., Godderis, L., Molenberghs, G., Meuris, C., Artesi, M., Durkin, K., Rahmouni, S., GREGOIRE, C., ... DARCIS, G. (14 June 2022). Evaluation of Screening Program and Phylogenetic Analysis of SARS-CoV-2 Infections among Hospital Healthcare Workers in Liège, Belgium. Viruses, 14 (6), 1302. doi:10.3390/v14061302  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Saegerman, C., Nguyet Diep, A., Renault, V., Donneau, A.-F., Stamatakis, L., Coppieters, W., Michel, F., Breuer, C., Dandoy, M., Ek, O., Gourzonès, C., Schyns, J., Goffin, E., Minner, F., Durkin, K., Artesi, M., Bours, V., Bureau, F., & Gillet, L. (2022). A 2-month field cohort study of SARS-CoV-2 in saliva of BNT162b2 vaccinated nursing home workers. Communications Medicine. doi:10.1038/s43856-021-00067-3  Peer reviewed Peer reviewed |
  | Thissen, R., ARTESI, M., Durkin, K., JOSSE, C., PALMEIRA, L., & BOURS, V. (24 December 2021). Comprehensive detection of homologous recombination deficiency by Nanopore sequencing [Poster presentation]. GIGA cancer day, Liège, Belgium. |
  | Durkin, K., Artesi, M., VAN DEN BROEKE, A., Bours, V., Hahaut, V., & Georges, M. (21 June 2021). Pooled crispr inverse pcr sequencing (pcip-seq): simultaneous sequencing of viral insertion points and the integrated viral genomes with long reads. |
  | Van Campenhout, C., De Mendonça, R., Alexiou, B., De Clercq, S., Racu, M.-L., Royer-Chardon, C., Rusu, S., Van Eycken, M., ARTESI, M., Durkin, K., Mardulyn, P., BOURS, V., Decaestecker, C., Remmelink, M., Salmon, I., & D'Haene, N. (2021). Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2) Genome Sequencing from Post-Mortem Formalin-Fixed, Paraffin-Embedded Lung Tissues. The Journal of Molecular Diagnostics. doi:10.1016/j.jmoldx.2021.05.016  Peer reviewed Peer reviewed |
Tang, L., Harland, C., Durkin, K., Artesi, M., Costa Monteiro Moreira, G., Karim, L., Deckers, M., Mullaart, E., Coppieters, W., Georges, M., & Charlier, C. (11 May 2021). Evidence for inter-individual variation of ERV de novo transposition rate in the male cattle germline [Paper presentation]. The Biology of Genomes, United States. |
  | Artesi, M.* , Hahaut, V.* , Cole, B.* , Lambrechts, L., Ashrafi, F., Marçais, A., Hermine, O., Griebel, P., Arsic, N., van der Meer, F., Burny, A., Bron, D., BIANCHI, E., DELVENNE, P., BOURS, V., Charlier, C., Georges, M., Vandekerckhove, L.* , Van den Broeke, A.* , & Durkin, K.*. (2021). PCIP-seq: simultaneous sequencing of integrated viral genomes and their integration sites with long reads. Genome Biology, 22, 97. doi:10.1186/s13059-021-02307-0  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi* These authors have contributed equally to this work. |
  | Marçais, A., Lhermite, L., Artesi, M., Laurent, C., Durkin, K., Hahaut, V., Rosewick, N., Suarez, F., Sibon, D., Cheminant, M., Avettand-Fenoël, V., Bruneau, J., Georges, M., Pique, C., Van den Broeke, A.* , Asnafi, V.* , & Hermine, O.*. (2021). Targeted deep sequencing reveals clonal and subclonal mutational signatures in Adult T-cell leukemia/lymphoma and defines an unfavorable indolent subtype. Leukemia, 35, 764-776. doi:10.1038/s41375-020-0900-3  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi* These authors have contributed equally to this work. |
  | Butera, Y., Mukantwari, E., Artesi, M., Umuringa, J. D. A., O'Toole, Á. N., Hill, V., Rooke, S., Hong, S. L., Dellicour, S., Majyambere, O., Bontems, S., Boujemla, B., Quick, J., Resende, P. C., Loman, N., Umumararungu, E., Kabanda, A., Murindahabi, M. M., Tuyisenge, P., ... Rujeni, N.*. (2021). Genomic sequencing of SARS-CoV-2 in Rwanda reveals the importance of incoming travelers on lineage diversity. Nature Communications, 12 (1), 5705. doi:10.1038/s41467-021-25985-7  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi* These authors have contributed equally to this work. |
  | Dellicour, S., Durkin, K., Hong, S., Vanmechelen, B., Marti-Carreras, J., Gill, M., Meex, C., Bontems, S., André, E., Gilbert, M., Walker, C., De Maio, N., Hadfield, J., Hayette, M.-P., Bours, V., Wawina-Bokalanga, T., Artesi, M., Baele, G., Maes, P., & Faria, N. R. (2021). A phylodynamic workflow to rapidly gain insights into the dispersal history and dynamics of SARS-CoV-2 lineages. Molecular Biology and Evolution. doi:10.1093/molbev/msaa284  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Maclot, F., Bontems, S., Meex, C., Artesi, M., Beckers, P., Bours, V., Durkin, K., & Hayette, M.-P. (2021). Development of a multiplex RT-qPCR using the drop out strategy to screen the SARS-CoV-2 South African 501Y.V2 variant. The Journal of infection, 83 (1), 19-e21. doi:10.1016/j.jinf.2021.04.035  Peer reviewed Peer reviewed |
  | Bollen, N., Artesi, M., Durkin, K., Hong, S. L., Potter, B., Boujemla, B., Vanmechelen, B., Martí-Carreras, J., Wawina-Bokalanga, T., Meex, C., Bontems, S., Hayette, M.-P., André, E., Maes, P., Bours, V., Baele, G., & Dellicour, S. (2021). Exploiting genomic surveillance to map the spatio-temporal dispersal of SARS-CoV-2 spike mutations in Belgium across 2020. Scientific Reports, 11 (1), 18580. doi:10.1038/s41598-021-97667-9  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Meuris, C., Kremer, C., Geerinck, A., Locquet, M., Bruyère, O., Defêche, J., Meex, C., Hayette, M.-P., Duchene, L., Dellot, P., Azarzar, S., Maréchal, N., Sauvage, A.-S., Frippiat, F., Giot, J.-B., Léonard, P., Fombellida, K., Moutschen, M., Durkin, K., ... Darcis, G. (2021). Transmission of SARS-CoV-2 After COVID-19 Screening and Mitigation Measures for Primary School Children Attending School in Liège, Belgium. JAMA Network Open, 4 (10), 2128757. doi:10.1001/jamanetworkopen.2021.28757  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
Tang, L., Harland, C., Durkin, K., Artesi, M., Costa Monteiro Moreira, G., Karim, L., Deckers, M., Mullaart, E., Coppieters, W., Georges, M., & Charlier, C. (06 October 2020). INTER-INDIVIDUAL VARIATION OF DE NOVO ERV TRANSPOSITION RATE IN THE CATTLE GERMLINE [Poster presentation]. Transposable Elements, United States. |
  | Artesi, M., Bontems, S., Göbbels, P., Franckh, M., Maes, P., BOREUX, R., MEEX, C., MELIN, P., Hayette, M.-P., Bours, V., & Durkin, K. (2020). A recurrent mutation at position 1 26,340 of SARS-CoV-2 is associated with failure of the E-gene qRT-PCR utilized in a commercial dual-target diagnostic assay. Journal of Clinical Microbiology. doi:10.1128/JCM.01598-20  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Artesi, M., Tamma, N., Deckers, M., Karim, L., Coppieters, W., Van den Broeke, A., Georges, M., Charlier, C., & Durkin, K. (2020). Colour-sidedness in Gloucester cattle is associated with a complex structural variant impacting regulatory elements downstream of KIT. Animal Genetics. doi:10.1111/age.12932  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Rosewick, N.* , Hahaut, V.* , Durkin, K., Artesi, M., Karpe, S., Wayet, J., Griebel, P., Arsic, N., Marçais, A., Hermine, O., Burny, A., Georges, M., & Van den Broeke, A. (2020). An Improved Sequencing-Based Bioinformatics Pipeline to Track the Distribution and Clonal Architecture of Proviral Integration Sites. Frontiers in Microbiology, 11, 587306. doi:10.3389/fmicb.2020.587306  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi* These authors have contributed equally to this work. |
Durkin, K. (27 March 2019). Simultaneous sequencing of retroviral insertion points and the associated provirus with Oxford Nanopore [Paper presentation]. Nanopore Day, Antwerp, Belgium. |
  | Artesi, M., Hahaut, V., Ashrafi, F., Marcais, A., Hermine, O., Griebel, P., Arsic, N., van der Meer, F., Burny, A., Bron, D., Charlier, C., Georges, M., Van den Broeke, A., & Durkin, K. (2019). Pooled CRISPR Inverse PCR sequencing (PCIP-seq): simultaneous sequencing of retroviral insertion points and the associated provirus in thousands of cells with long reads 558130. ORBi-University of Liège. https://orbi.uliege.be/handle/2268/237583. doi:10.1101/558130 |
  | Galli, V., Nixon, C. C., Strbo, N., Artesi, M., de Castro-Amarante, M. F., McKinnon, K., Fujikawa, D., Omsland, M., Washington-Parks, R., Romero, L., Caruso, B., Durkin, K., Brown, S., Karim, B., Vaccari, M., Jacobson, S., Zack, J. A., Van den Broeke, A., Pise-Masison, C., ... Silvestri, G. (2019). Essential Role of Human T Cell Leukemia Virus Type 1 orf-I in Lethal Proliferation of CD4+ Cells in Humanized Mice. Journal of Virology, 93 (19 e00565-19). doi:10.1128/JVI.00565-19  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Georges, M., Van den Broeke, A., Durkin, K., Artesi, M., Hahaut, V., & Rosewick, N. (11 October 2018). MONITORING METHOD FOR ADULT T-CELL LEUKEMIA/LYMPHOMA (ATL). |
  | Hahaut, V., Rosewick, N., Artesi, M., Durkin, K., Burny, A., Bron, D., Georges, M., & Van den Broeke, A. (02 February 2018). Improving the bioinformatics analysis of HTS clonality data in virus-induced leukemia [Poster presentation]. Télévie Cancer Seminar 2018. |
Durkin, K. (2018). Farm to variant, Using domestic animal genomics to explore basic biological questions. |
Durkin, K., Artesi, M., Hahaut, V., Rosewick, N., Griebel, P., Arsic, N., Burny, A., Georges, M., & Van den Broeke, A. (21 September 2017). The Landscape And Evolution Of Somatic Mutations In Bovine Leukemia Virus Induced Tumors [Paper presentation]. Animal Genetics & Diseases, Wellcome Genome Campus, Hinxton, Cambridge, United Kingdom. |
Durkin, K., Artesi, M., Hahaut, V., Rosewick, R., Griebel, P., Arsic, N., Burny, A., Georges, M., & Van den Broeke, A. (July 2017). Somatic Structural And Numerical Aberrations In Bovine Leukemia Virus Induced Tumors [Poster presentation]. 36th International Society for Animal Genetics Conference, Dublin, Ireland. |
  | Rosewick, N.* , Durkin, K.* , Artesi, M.* , Marçais, A., Hahaut, V., Griebel, P., Avettand-Fenoel, V., Burny, A., Charlier, C., Hermine, O., Georges, M., & Van den Broeke, A. (23 May 2017). Cis-perturbation of cancer drivers by the HTLV-1/BLV proviruses is an early determinant of leukemogenesis. Nature Communications, 8. doi:10.1038/ncomms15264  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi* These authors have contributed equally to this work. |
  | Hahaut, V., Artesi, M., Durkin, K., Rosewick, N., Griebel, P., Natasa, A., Burny, A., Georges, M., & Van den Broeke, A. (08 March 2017). Investigating non-coding viral transcripts in Bovine Leukemia Virus induced leukemia [Poster presentation]. 18th International Conference on Human Retrovirology: HTLV and Related Viruses, Tokyo, Japan. |
Durkin, K., Artesi, M., Hahaut, V., Rosewick, N., Griebel, P., Arsic, N., Burny, A., Georges, M., & Van den Broeke, A. (March 2017). Structural And Numerical Somatic Changes In BLV Induced Tumors [Poster presentation]. 18th International Conference on Human Retorovirology HTLV and Related Viruses, Tokyo, Japan. |
  | Artesi, M., Marcais, A., Durkin, K., Rosewick, N., Hahaut, V., Suarez, F., Trinquand, A., Lhermitte, L., Asnafi, V., Avettand-Fenoel, V., Burny, A., Georges, M., Hermine, O., & Van den Broeke, A. (2017). Monitoring molecular response in adult T-cell leukemia by high-throughput sequencing analysis of HTLV-1 clonality. Leukemia, 31 (11), 2532-2535. doi:10.1038/leu.2017.260  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
Marçais, A., Artesi, M., Durkin, K., Hahaut, V., Suarez, F., Sibon, D., Cheminand, M., Delarue, R., Lhermitte, L., Asnafi, V., Avettand Fenoel, V., Georges, M., Van den Broeke, A., & Hermine, O. (2017). Improved high throughput DNA-seq based mapping of HTLV-1 integration sites: a tool to define response to treatment, ATL prognosis and guide therapeutic strategies [Paper presentation]. 18th international conference on human retrovirology - HTLV and related viruses, Tokyo, Japan. |
Durkin, K., Rosewick, N., Artesi, M., Hahaut, V., Griebel, P., Arsic, N., Burny, A., Georges, M., & Van den Broeke, A. (21 May 2016). Identification and characterization of novel bovine leukemia virus (BLV) antisense transcripts in leukemic and pre-leukemic clones [Paper presentation]. HTLV European Research Network Meeting 2016, Bucharest, Romania. |
  | Durkin, K., Rosewick, N., Artesi, M., Hahaut, V., Griebel, P., Arsic, N., Burny, A., Georges, M., & Van den Broeke, A. (2016). Characterization of novel Bovine Leukemia Virus (BLV) antisense transcripts by deep sequencing reveals constitutive expression in tumors and transcriptional interaction with viral microRNAs. Retrovirology, 13 (1), 33. doi:10.1186/s12977-016-0267-8  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
Artesi, M., Durkin, K., Rosewick, N., Georges, M., & Van den Broeke, A. (2016). Improving the methodology for high throughput mapping of proviral integration sites [Paper presentation]. The HTLV european research network meeting, Bucharest, Romania. |
  | Durkin, K., Artesi, M., Rosewick, N., Marçais, A., Hermine, O., Georges, M., & Van den Broeke, A. (28 August 2015). Improving the methodology for the detection of proviral integration sites in the host genome via high throughput sequencing. Retrovirology, 12 (1).  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Rosewick, N., Durkin, K., Marçais, A., Artesi, M., Hermine, O., Charlier, C., Georges, M., & Van den Broeke, A. (28 August 2015). HTLV-1/BLV antisense-RNA dependent host gene perturbation in pre-leukemic and leukemic clones. Retrovirology, 12 (1).  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
Durkin, K., Artesi, M., Rosewick, N., Georges, M., & Van den Broeke, A. (May 2015). Improving proviral integration site detection with high throughput sequencing [Poster presentation]. The biology of genomes, Cold Spring Harbor, United States - New York. |
Durkin, K., Rosewick, N., Momont, M., Thys, W., Burny, A., Georges, M., & Van den Broeke, A. (June 2014). Identification of two noncoding antisense transcripts in BLV and their interaction with the BLV encoded miRNAs [Paper presentation]. HERN meeting, Rome, Italy. |
Rosewick, N., Durkin, K., Thys, W., Marçais, A., Hermine, O., Burny, A., Georges, M., & Van den Broeke, A. (June 2014). Exploring the Deltaretrovirus Tumor Transcriptome: Lessons from RNA-Seq [Paper presentation]. HERN meeting, Rome, Italy. |
Durkin, K., Rosewick, N., Momont, M., Thys, W., Burny, A., Georges, M., & Van den Broeke, A. (February 2014). Identification of two antisense transcripts in BLV and their interaction with BLV-encoded microRNAs [Poster presentation]. Keystone meeting Non-coding RNAs: towards mechanisms, Santa Fe, United States. |
  | Durkin, K., Rosewick, N., Momont, M., Thys, W., Burny, A., Georges, M., & Van den Broeke, A. (07 January 2014). Elucidating the role of Bovine Leukemia Virus encoded micro-RNAs. Retrovirology, 11 (1), 62.  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
Durkin, K. (28 January 2013). Deep sequencing reveals abundant non-canonical retroviral microRNAs in B-cell leukemia/lymphoma [Paper presentation]. GIGA-Day. |
  | Rosewick, N., Momont, M., Durkin, K., Takeda, H., Caiment, F., Cleuter, Y., Vernin, C., Mortreux, F., Wattel, E., Burny, A., Georges, M., & Van den Broeke, A. (2013). Deep sequencing reveals abundant non-canonical retroviral microRNAs in B-cell leukemia/lymphoma. Proceedings of the National Academy of Sciences of the United States of America.  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Dupuis, M.-C., Zhang, Z., Durkin, K., Charlier, C., Lekeux, P., & Georges, M. (2013). Detection of copy number variants in the horse genome and examination of their association with recurrent laryngeal neuropathy. Animal Genetics. doi:10.1111/j.1365-2052.2012.02373.x  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
Durkin, K. (04 May 2012). Serial translocation via circular intermediates underlies color-sidedness in cattle [Paper presentation]. GIGA-Day. |
  | Durkin, K., Coppieters, W., Drogemuller, C., Ahariz, N., Cambisano, N., Druet, T., Fasquelle, C., Haile, A., Horin, P., Huang, L., Kamatani, Y., Karim, L., Lathrop, M., Moser, S., Oldenbroek, K., Rieder, S., Sartelet, A., Solkner, J., Stalhammar, H., ... Charlier, C. (02 February 2012). Serial translocation by means of circular intermediates underlies colour sidedness in cattle. Nature, 482 (7383), 81-4. doi:10.1038/nature10757  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Niku, M., Liljavirta, J., Durkin, K., Schroderus, E., & Iivanainen, A. (2012). The bovine genomic DNA sequence data reveal three IGHV subgroups, only one of which is functionally expressed. Developmental and Comparative Immunology, 37, 457-461. doi:10.1016/j.dci.2012.02.006  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
Durkin, K., Cambisano, N., Ahariz, N., Fasquelle, C., Karim, L., Sartelet, A., Drogemüller, C., Leeb, T., Coppieters, W., Georges, M., & Charlier, C. (May 2011). Molecular dissection of the color-sided pehnotype in cattle reveals a novel mechanism of chromosome evolution involving circular shuttling intermediates [Poster presentation]. The Biology Of Genomes, Cold Spring Harbor, United States - New York. |
  | Durkin, K., Cambisano, N., Ahariz, N., Fasquelle, C., Karim, L., Sartelet, A., Baurain, D., Drogemüller, C., Leeb, T., Coppieters, W., Georges, M., & Charlier, C. (May 2011). Molecular dissection of the color-sided phenotype in cattle reveals a novel mechanism of chromosome evolution involving circular shuttling intermediates. Chromosome Research, 19 (S1), 18.  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
Durkin, K., Raudsepp, T., & Chowdhary, B. P. (2011). Cytogenetic Evaluation of the Stallion. In A. O. McKinnon, E. L. Squires, W. E. Vaala, ... D. D. Varner (Eds.), Equine Reproduction (2nd Edition, pp. 1462-1468). Wiley-Blackwell.  Peer reviewed Peer reviewed |
  | Raudsepp, T., Durkin, K., Lear, T. L., Das, P. J., Avila, F., Kachroo, P., & Chowdhary, B. P. (2010). Molecular heterogeneity of XY sex reversal in horses. Animal Genetics, 41, 41-52. doi:10.1111/j.1365-2052.2010.02101.x  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Durkin, K. (2009). Mapping Athletic Performance Related Genes in the Equine Genome and a Genome Scan for Superior Athletic Performance in the Thoroughbred [Doctoral thesis, Texas A&M University]. ORBi-University of Liège. https://orbi.uliege.be/handle/2268/138232 |
Durkin, K., Raudsepp, T., Skow, L. C., & Chowdhary, B. P. (July 2008). A genome scan for athletic performance in the thoroughbred [Poster presentation]. XXXI Conference of the International Society for Animal Genetics (ISAG), Amsterdam, Netherlands. |
Durkin, K., Raudsepp, T., & Chowdhary, B. P. (2008). Precise demarcation of the breakpoint in a thoroughbred stallion carrying a ECA5 and ECA16 Translocation. In Proceedings of Plant & Animal Genome XVI.  Peer reviewed Peer reviewed |
  | Raudsepp, T., Gustafson-Seabury, A., Durkin, K., Wagner, M. L., Goh, G., Seabury, C., Brinkmeyer-Langford, C., Lee, E. J., Agarwala, R., Stallknecht-Rice, E., Schäffer, A. A., Skow, L., Tozaki, T., Yasue, H., Penedo, M. C. T., Lyons, L. A., Khazanehdari, K. A., Binns, M. M., MacLeod, J. N., ... Chowdhary, B. P. (2008). A 4,103 marker integrated physical and comparative map of the horse genome. Cytogenetic and Genome Research, 122, 28-36. doi:10.1159/000151313  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Brito, L. F. C., Sertich, P. L., Durkin, K., Chowdhary, B. P., Turner, R. M., Greene, L. M., & McDonnell, S. (2008). Autosomic 27 Trisomy in a Standardbred Colt. Journal of Equine Veterinary Science, 28 (7), 431-436. doi:10.1016/j.jevs.2008.06.003  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Goh, G., Raudsepp, T., Durkin, K., Wagner, M. L., Schäffer, A. A., Agarwala, R., Tozaki, T., Mickelson, J. R., & Chowdhary, B. P. (2007). High-resolution gene maps of horse chromosomes 14 and 21: Additional insights into evolution and rearrangements of HSA5 homologs in mammals. Genomics, 89, 89-122. doi:10.1016/j.ygeno.2006.06.012  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
Durkin, K., Raudsepp, T., & Chowdhary, B. P. (2006). Cytogenomic analyses for fertility and sex determination defects in the horse. In Proceedings of the 33rd Annual Meeting of the Texas Genetics Society.  Peer reviewed Peer reviewed |
Durkin, K., Raudsepp, T., Skow, L. C., & Chowdhary, B. P. (January 2006). A map of performance related genes in the horse [Poster presentation]. Plant & Animal Genome XIV Conference, San Diego, United States - California. |
  | Perrocheau, M., Chadi, S., Mata, X., Decaunes, P., Raudsepp, T., Durkin, K., Incarnato, D., Iannuzzi, L., Lear, T. L., Hirota, K., Hasegawa, T., Zhu, B., de Jong, P., Cribiu, E. P., Chowdhary, B. P., & Guerin, G. (2006). Construction of a medium-density horse gene map. Animal Genetics. doi:10.1111/j.1365-2052.2005.01401.x  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
  | Durkin, K., Raudsepp, T., & Chowdhary, B. P. (2006). Characterization of two structural aberrations in the horse by FISH with BAC clones. Chromosome Research, 14, 213-220.  Peer Reviewed verified by ORBi Peer Reviewed verified by ORBi |
Durkin, K., Raudsepp, T., & Chowdhary, B. P. (2005). Chromosome aberrations and equine fertility. In Proceedings of the 32nd Annual Meeting of the Texas Genetics Society.  Peer reviewed Peer reviewed |
Durkin, K., Raudsepp, T., Skow, L. C., & Chowdhary, B. P. (January 2005). Analyzing athletic performance in thoroughbred horses [Poster presentation]. Plant & Animal Genome XIII, San Diego, United States - California. |